Inhalt
Reverse Clinical Engineering®
Using patient-derived 3D tumor models as in vitro diagnostics for modern precision oncology
ASC Oncology has developed Reverse Clinical Engineering® (RCE), a test procedure to determine in individual patients with malignant solid tumors which anticancer treatments are most likely to be effective and which are not. The RCE is an in vitro diagnostic test based on patient-derived 3D (PD3D®) cell cultures, so-called tumor organoids, obtained from a patient’s vital tumor sample. The test provides the treating physician with additional information, thus making it easier for both the physician and the patient to decide on an optimal therapy. This enables a personalized therapy with presumably higher likelihood of survival and a better quality of life to be selected as early as possible, and treatments that are probably ineffective, including potential undesired side effects, to be avoided.
The Reverse Clinical Engineering® test procedure, a form of personalized medicine, enables therapy decisions to be made based on individual in vitro data.
Medical need
The Reverse Clinical Engineering® test procedure is a valuable complementary diagnostic tool for deciding on the optimal therapy
Tumors exhibit significant heterogeneity, which presents a challenge for treatment (Burrell et al., 2013): while many tumors respond to recommended standard treatment regimens, there is a substantial proportion of non-responders for whom established therapeutic concepts do not have the desired effect. By providing complementary data for medical treatment decisions, RCE testing helps physicians to offer their patients the most appropriate therapy with the highest chances of success.
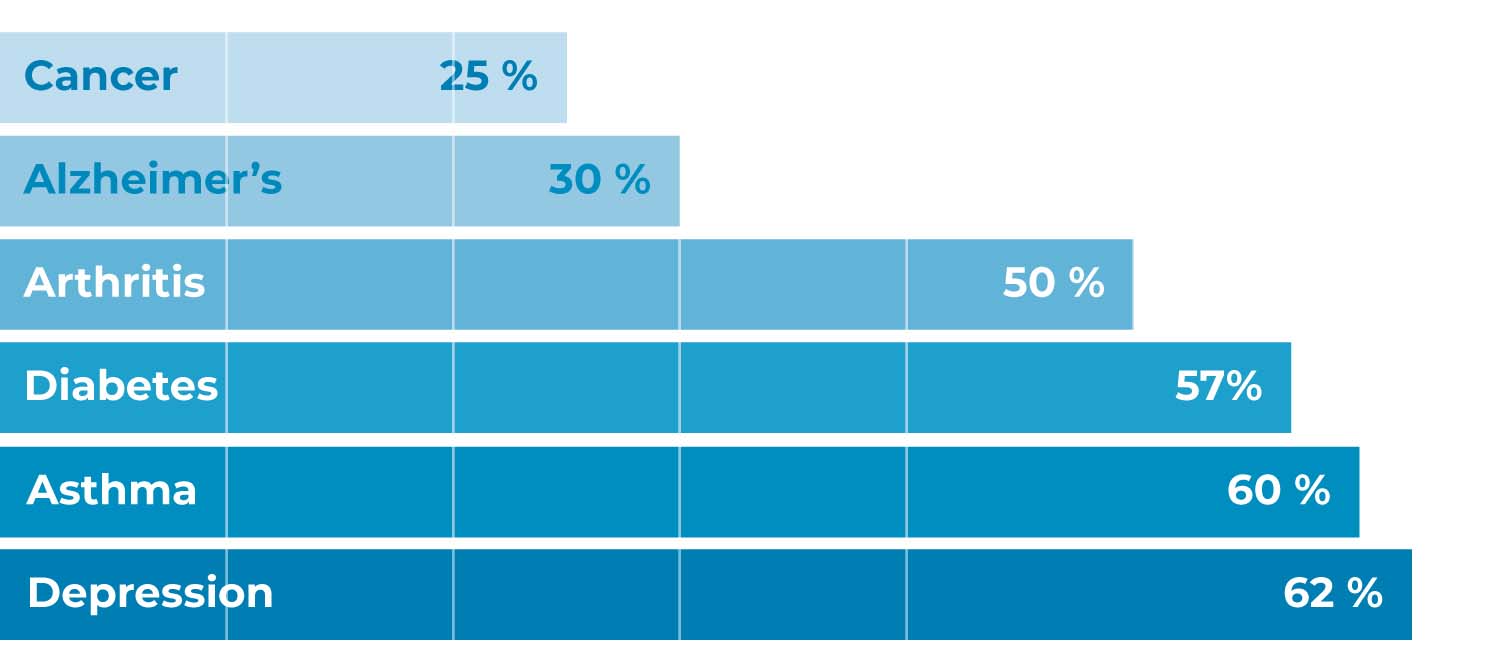
Percentage of patients for whom an administered drug is effective, for selected diseases (Statista, 2021).
Reverse Clinical Engineering® + clinical trial data = more insights
The test procedure is an important supplement to randomized controlled trials (RCTs). Similarly, to an n = 1 study, it can determine the efficacy of drug therapies for each individual patient prior to commencing therapy. In contrast, RCTs can evaluate a therapy’s overall efficacy and safety. However, as individual responses can only be derived to a limited extent from the average values obtained in large cohort studies, the informative value for the individual patient is limited.

King’s College London & University College London Cancer Institute
On the way to improved individual therapy decisions with Reverse Clinical Engineering®
Personalized therapy concepts and stratification according to molecular tumor subtypes are becoming increasingly common in clinical practice. However, precision medicine is still in its infancy: studies have found that sequencing identifies therapeutically actionable driver mutations in less than half of analyzed patients, and only 0.8-3% of patients subsequently benefit from matched targeted therapy (Tannock & Hickman, 2019).
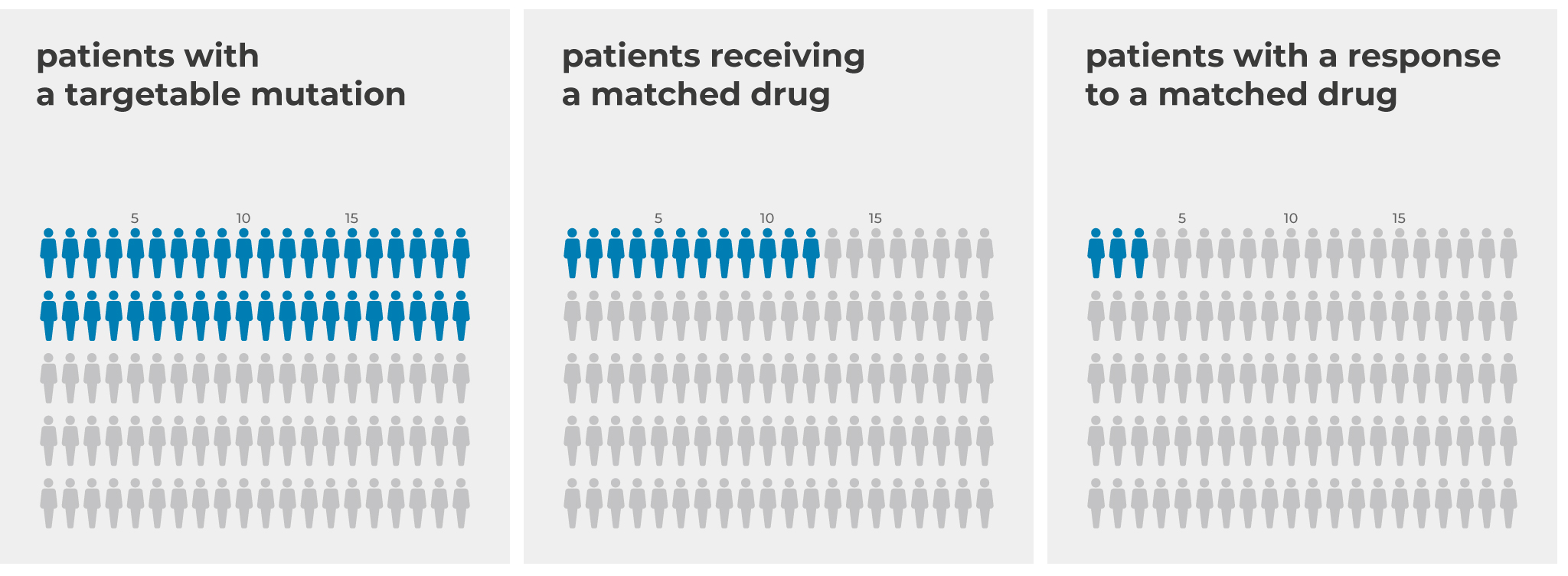
Only a small proportion of patients who underwent genetic testing and were subsequently treated with mutation-targeted therapy were able to benefit from targeted treatment (Tannock & Hickman, 2019).
There is growing scientific evidence that tumors exhibit heterogeneity and/ or plasticity that extends beyond the genetic level (Brock & Huang, 2017). Cell connectivity and the microenvironment seem to influence how the tumor responds to treatment and whether resistance develops (Huang, 2021). While the impact of plasticity on the tumor’s therapy response cannot yet be adequately mapped by genome sequencing or other forms of biomarker testing, some of it can be replicated in patient-derived 3D tumor models (Walter et al., 2021, Yao 2020, Yang et al., 2022) – which is why the RCE is a valuable addition to standard diagnostics.

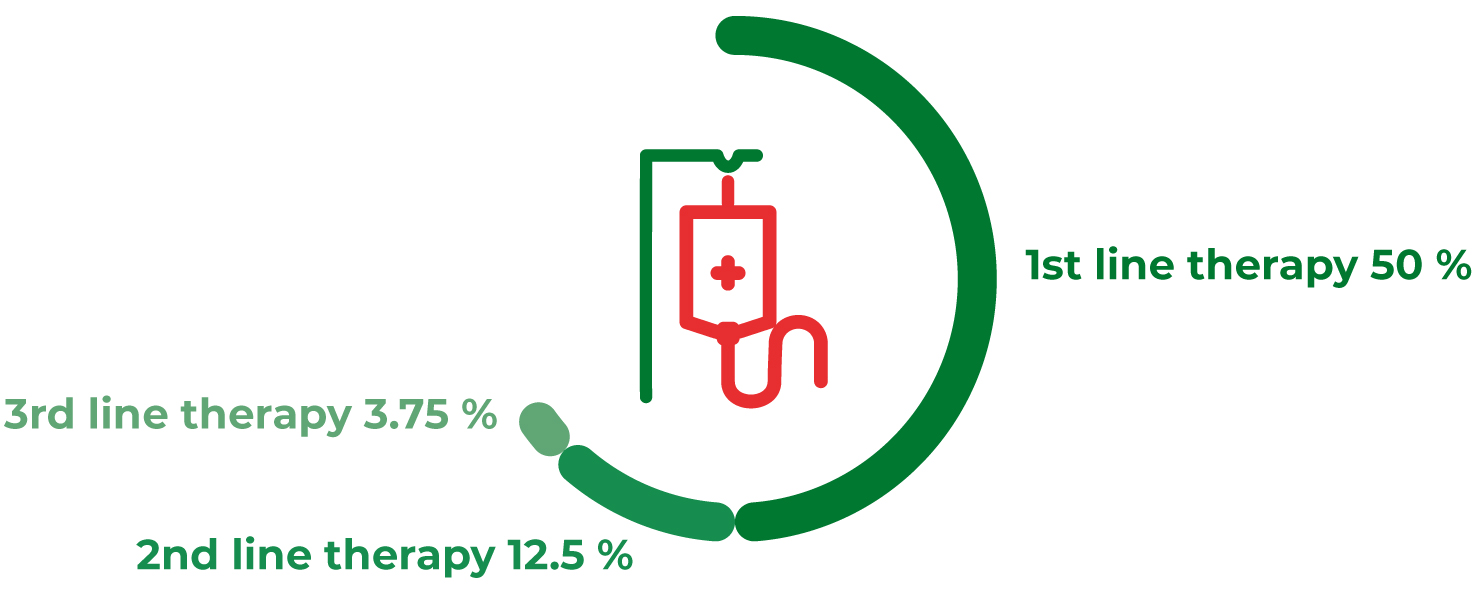
Reverse Clinical Engineering® to reduce the risk of further lines of therapy
The patient’s prognosis worsens with each additional line of therapy, because the likelihood of a therapy response can decrease considerably (Kessous et al., 2020). RCE testing helps to identify the best possible treatment option early on, which can minimize the risk of ineffective treatment and subsequent further lines of therapy. In addition, the procedure can help identify the best possible therapeutic option in a timely manner when guideline recommendations are only available with a low level of evidence, as in certain rare tumors or advanced tumors for which there are no further curative options.
How the test procedure works
The RCE test procedure is based on PD3D® tumor models, so-called organoids. Together with partners from pharmaceutical research, our scientists have developed an automated platform which enables the screening of organoids in a high-throughput, parallelized and standardized manner (Boehnke et al., 2016). The procedure has replaced the use of the NCI60 panel for pre-clinical drug testing at the National Cancer Institute (NCI) and has been selected by the NCI as a “best practice approach” (Ledford, 2016, Boehnke et al., 2016, NCI, 2019).

King’s College London & University College London Cancer Institute
From a fresh tissue sample to the patient-derived 3D tumor model
PD3D® tumor models can be generated from fresh tissue samples, such as resects or biopsies. The sample is, first, mechanochemically processed in the laboratory and living tumor cells are then cultivated in a matrix. The matrix resembles the human extracellular basement membrane and thus mimics the tumor’s complex environment. In addition, the matrix forms a hydrogel that allows the cells to grow three-dimensionally. Each individual tumor entity is cultured in a specifically optimized medium that favors the tumor cells’ growth and limits the proliferation of other normal non-cancerous cells in the sample. Under these conditions, the tumor cells develop into organoids, which are composed of organ-specific cell types and spatially organized in a manner similar to the tumor’s parent organ.
The organoids exhibit genomic, molecular, and phenotypic characteristics of the patient’s tumor that can be stably maintained over multiple passages in long-term cultures (Sachs et al.,2018). As accurate and clinically relevant copies, they can also replicate different subtypes and the intratumoral heterogeneity of the patient’s tumor (Kopper et al., 2019, Sharick et al., 2020).
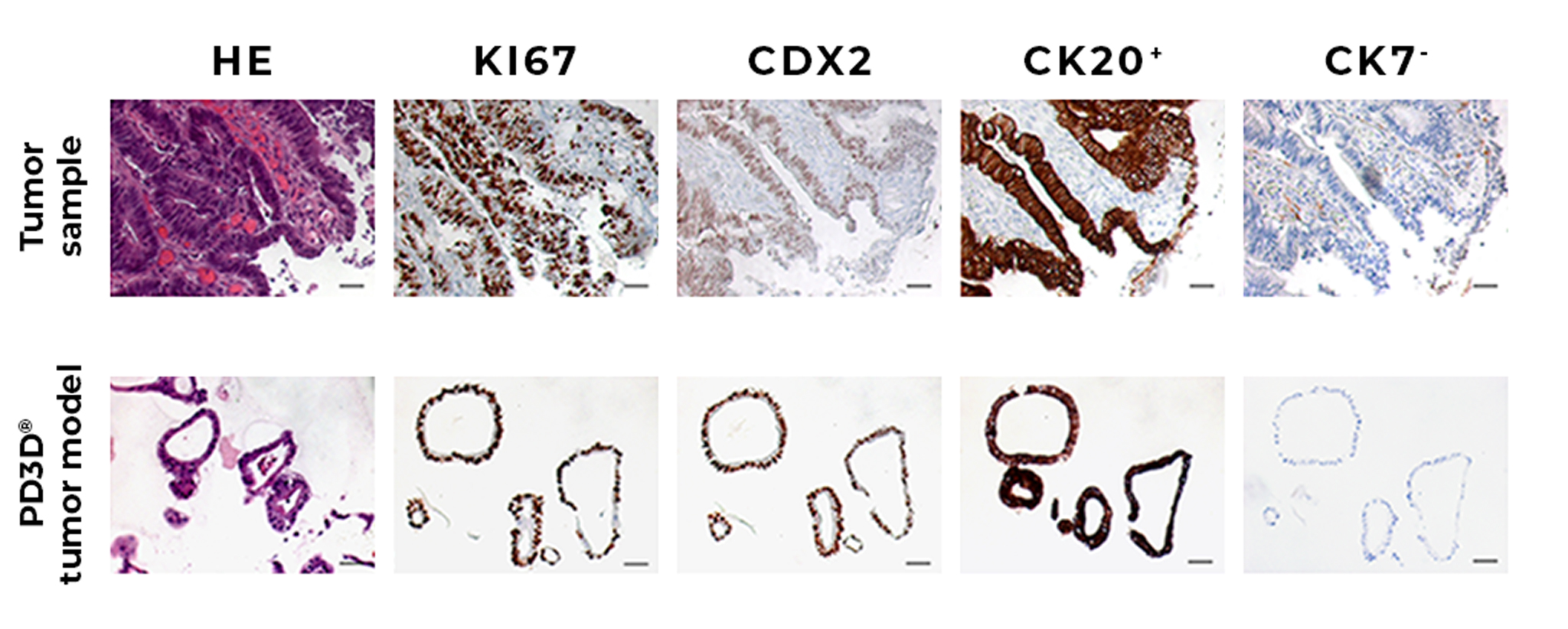
Comparison between immunohistochemical staining of a primary tumor and the derived PD3D® tumor model stained for hematoxylin-eosin (HE), CDX2, CK20+ and CK7-. Ki67 expression marks actively proliferating cells in both original tissue and tumor model (Schütte et al., 2017 Suppl.).
From testing to the chemosensitivity profile
Once PD3D® models have been created and amplified, the drugs in question can be tested. For this purpose, the organoids are cultured in 96- or 384-well plates, treated with the drugs in an automated and parallelized manner, and the cell viability is then measured and evaluated.
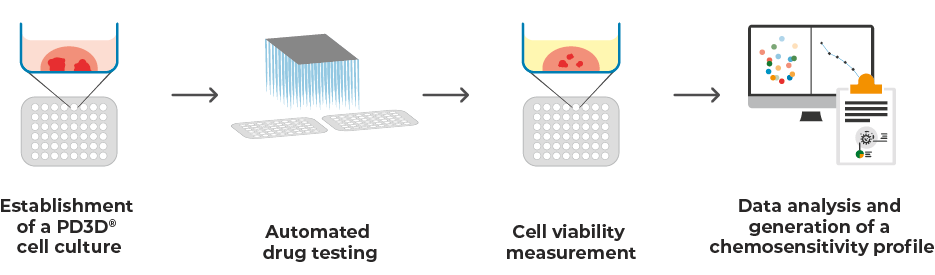
Workflow of Reverse Clinical Engineering®
To obtain clinically relevant results, the active substances are assayed at different concentrations, which are based on the clinically measurable maximum concentration (Cmax) of the respective active substance in blood plasma. Cell viability is determined using an ATP-dependent luminescence assay. This quantifies the ATP content of the treated organoids, which is directly proportional to the number of metabolically active viable cells (Crouch et al., 1993).
The data is used to determine the IC50 value, which indicates the drug concentration at which growth of the tumor cells is half-maximally inhibited. If the IC50 value is above the Cmax value (IC50 > Cmax), it is unlikely that therapy will result in an effective tumor response in the patient. A lower IC50, below Cmax (IC50 < Cmax), is more likely to inhibit tumor cell growth and result in a treatment response.
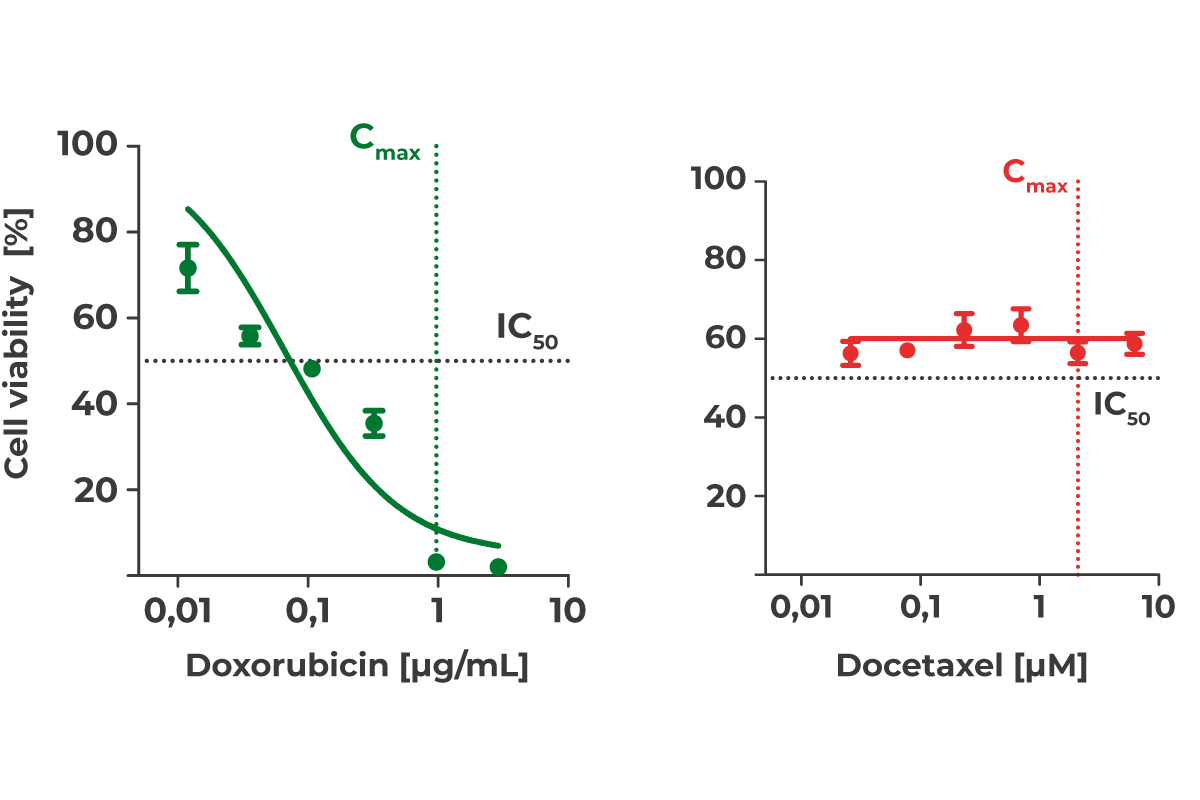
Dose-response curves for doxorubicin (green, IC50 < Cmax) and docetaxel (red, IC50 > Cmax).
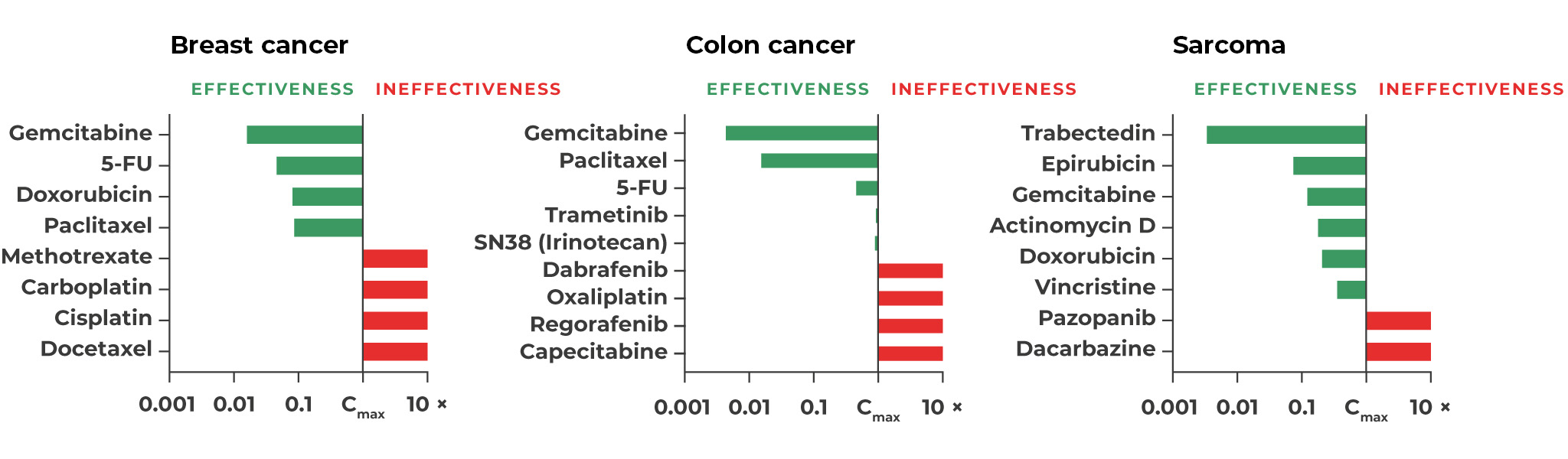
Response of different PD3D® tumor models to selected substances oriented to the clinically maximal blood plasma level (green: IC50 < Cmax, red: IC50 > Cmax).
Finally, within on average of 28 days after receiving the sample, we prepare a report containing a detailed evaluation and graphical presentation of the data as well as clear recommendations for the subsequent therapy decision. NB: The results should always be interpreted in the context of other medical data and understood as a valuable, complementary aid in decision-making and therapy planning with the patient.
Performance
The use of tumor organoids for improved personalized therapy has been demonstrated in international co-clinical studies. The procedure has also been recommended by the NCI as a “best practice approach” for drug testing since 2017 and is already established in the preclinical setting (Boehnke et al., 2016, NCI, 2019). ASC Oncology is preparing a multicenter international phase IV study during first-line therapy of difficult-to-treat tumor diseases with the Medical University of Innsbruck as the lead investigational site.
Medical Statistics
A meta-analysis of 17 co-clinical studies (in colorectal cancer, rectal cancer, gastric cancer, pancreatic cancer, ovarian cancer, and breast cancer; Wensink et al., 2021) found the following characteristics for the test procedure using tumor organoids:
Diagnostic sensitivity: 81% (95% CI: 0.69 – 0.89)
Diagnostic specificity: 74% (95% CI: 0.64 – 0.82)
For individual tumor entities, co-clinical studies revealed the following values:
Metastatic cancer of the gastrointestinal tract (n = 15)
Diagnostic sensitivity: 100%
Diagnostic specificity: 93%
Positive predictive value: 88%
Negative predictive value: 100%
Squamous cell carcinoma of the head and neck (n = 7)
Diagnostic sensitivity: 75%
Diagnostic specificity: 100%
Positive predictive value: 100%
Negative predictive value: 75%
Advanced colorectal cancer (n = 17)
Diagnostic sensitivity: 100%
Diagnostic specificity: 86%
Positive predictive value: 92%
Negative predictive value: 100%
Pancreatic cancer (n = 9)
Diagnostic sensitivity: 100%
Diagnostic specificity: 67%
Positive predictive value: 86%
Negative predictive value: 100%
Advanced lung cancer (n = 54)
Diagnostic sensitivity: 84%
Diagnostic specificity: 83%
Positive predictive value: 81%
Negative predictive value: 86%
Efficacy – higher chance of improved treatment response
Significantly prolonged progression-free survival (PFS) following patient-derived tumor cell assays
In patients with advanced colorectal cancer, the efficacy of adjuvant chemotherapy after tumor resection could be predicted using organoid-based assays. Accordingly, patients whose tumor organoids in the test were sensitive to chemotherapy had a significantly longer PFS of 16.0 months than patients whose tumor organoids in the test were resistant to chemotherapy and who had a shorter PFS of 9.0 months (p < 0,001; Wang et al., 2023).
Organoid-based assays were used in another study in patients with metastatic colorectal cancer to assess response to FOLFOX therapy. Patients whose tumor organoids were resistant to FOLFOX in the assay had a more than 4-fold higher risk of progression or death than patients whose tumor organoids were sensitive to therapy in the assay (HR: 4.3; 95% confidence interval: 2.6-7.2; p < 0.001; Tang et al., 2023). Median PFS was 8 months in the organoid-resistant patient group and 11 months in the organoid-sensitive group.
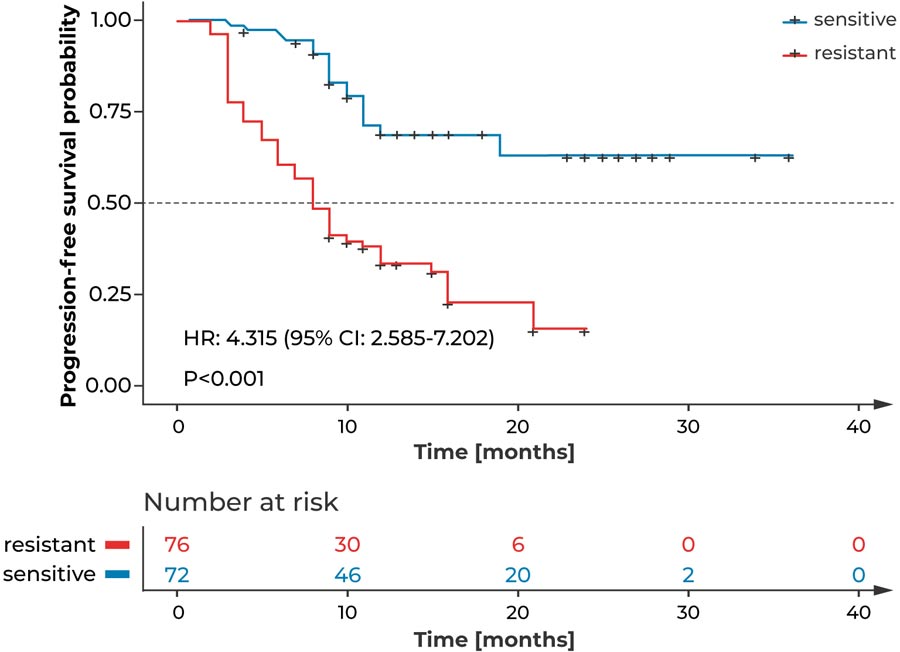
Kaplan-Meier curves show that patients with metastatic colorectal cancer who received FOLFOX therapy had a significantly higher risk of disease progression or death if their organoids had previously tested resistant to FOLFOX, compared with patients whose organoids were sensitive to FOLFOX in the test (Tang et al., 2023). CI = confidence interval, FOLFOX = folinic acid, fluorouracil, oxaliplatin, HR = hazard ratio
In patients with lung cancer, the use of patient-derived tumor cell testing has significantly prolonged the PFS by more than twofold, from 8.5 to 18 months (p < 0,05; Chen et al., 2018).
Prolonged relapse-free survival (RFS) in predicted responders according to tests with patient-derived tumor organoids
Patients with pancreatic cancer who were classified as responders to adjuvant therapy based on in vitro testing of individual tumor organoids had an RFS >12 months after surgical resection, whereas disease recurred within one year on adjuvant therapy in patients classified as non-responders based on in vitro data (Sharick et al., 2020).
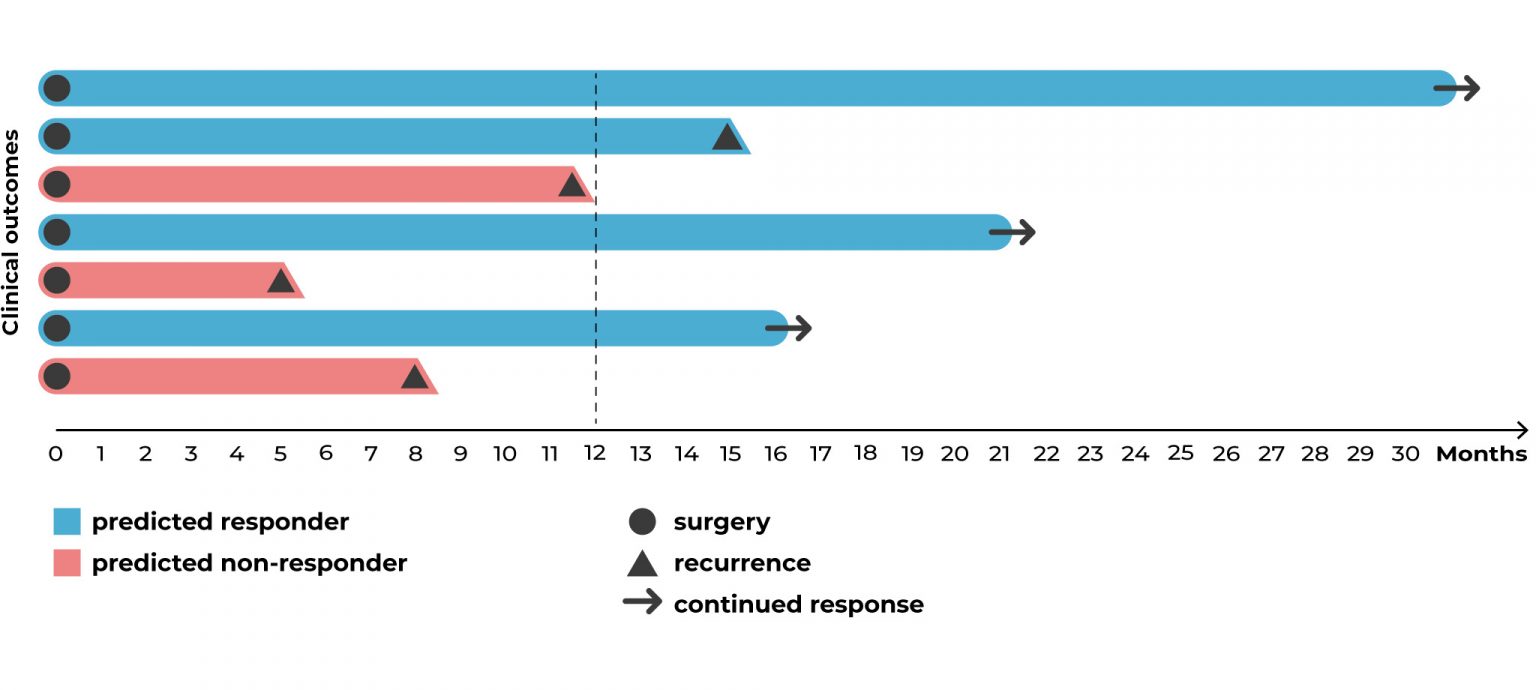
Patients with pancreatic cancer had prolonged recurrence-free survival after receiving therapy classified as effective by the in vitro testing of individual tumor organoids compared with patients receiving therapy classified as ineffective by in vitro testing (Sharick et al., 2020). The arrows indicate that patients were still alive without recurrence at the time the data was published.
Quality of life – Improved patient satisfaction
Increased adherence and avoidance of potential undesired side effects from less promising therapies through precision medicine
Tailored treatment plans with minimal side effects and patient participation in treatment decisions can benefit adherence, trust, quality of care, and cancer treatment outcomes (Shingler et al., 2014, Kehl et al., 2015).
Added value for the healthcare system
Avoiding presumably ineffective therapies can preserve the health care system’s limited resources, and cost-intensive treatments can be used more effectively. This can lead to relevant cost savings and free personnel capacities.
Requirements and limitations
Suitable specimen for the test procedure
- fresh tissue from malignant solid tumors of any stage (primary tumor or metastases)
- obtained by resection or biopsy
- before or during ongoing treatment (the patient may not have had anticancer therapy within two weeks of specimen collection)
Unsuitable specimen for the test procedure
- specimen from leukemias or lymphomas
- fixed tissue
- tissue samples with insufficient number of vital cells
Substances suitable for the test procedure
- cytostatics
- targeted therapies
- antibodies
- drugs used in off-label ways
- drugs currently under clinical investigation
- experimental, alternative, or naturopathic medicines
- as monotherapy or combination therapy
Substances unsuitable for the test procedure
- substances that must be metabolized and whose active metabolite is not available in a chemically stable form
- substances that have an indirect effect on the tumor and exert their effect on the tumor exclusively via other cell types
FAQ
What distinguishes the Reverse Clinical Engineering® test procedure from other tests?
Using the PD3D® tumor models, the cancer patient’s individual therapy response can be evaluated in vitro in an n=1 study prior to commencing treatment. Using the data in addition to other clinical parameters allows optimal decisions to be made for a predictively effective therapy option.
Patient-derived xenograft models in mice also allow an individual patient’s tumor response to be predicted. These models, however, are relatively time consuming: they take significantly longer to provide results, about 4-6 months compared to an average of 28 days using the Reverse Clinical Engineering® test procedure.
Moreover, sequencing technologies can reveal a tumor’s mutational status of, thus being able to provide clues to potential targeted therapy. However, the ability to predict whether the tumor will actually respond to such therapy prior to treatment is limited as, without individual data, this requires extrapolation from the averages of large cohort studies.
How long does the Reverse Clinical Engineering® test procedure take?
On average, from receiving the sample to issuing the final report, Reverse Clinical Engineering® testing takes 28 days. The procedure’s duration may vary depending on tumor entity, sample quality and sample quantity, as these factors influence the time required to grow and amplify the PD3D® tumor models.
What results does the Reverse Clinical Engineering® test procedure report from ASC Oncology provide?
The report provides a tabular overview of the experimentally determined IC50 values of all tested substances as well as a specific recommendation for the subsequent therapy decision. The data are additionally graphed to better illustrate the individual sensitivity profile. Furthermore, the report includes a detailed explanation of the method as well as definitions of the most important terms. Please take a look at an exemplary report.
Further reading
Boehnke K, Iversen PW, Schumacher D, Lallena MJ, Haro R, Amat J, Haybaeck J, Liebs S, Lange M, Schäfer R, Regenbrecht CRA, Reinhard C, Valesco A: Assay Establishment and Validation of a High-Throughput Screening Platform for Three-Dimensional Patient-Derived Colon Cancer Organoid Cultures. J Biomal Screen 21, 931-41 (2016).
This work describes the establishment of our platform for automated and parallelized drug testing using organoids.
Gaebler M, Silvestri A, Haybaeck J, Reichardt P, Lowery CD, Stancato LF, Zybarth G, Regenbrecht CRA: Three-Dimensional Patient-Derived In Vitro Sarcoma Models: Promising Tools for Improving Clinical Tumor Management. Front Oncol 11, 7 (2017)
This review provides a comprehensive overview of how organoid models can be used to better understand the molecular biology of sarcomas.
The first study to create a colorectal cancer biobank based on patient samples and preclinical models, and to provide a drug sensitivity data archive that can be used to improve drug development and understanding of colorectal cancer biology.
Schumacher D, Andrieux G, Boehnke K, Keil M, Silvestri A, Silvestrov M, Keilholz U, Haybaeck J, Erdmann G, Sachse C, Templin M, Hoffmann J, Boerries M, Schäfer R, Regenbrecht CRA: Heterogeneous pathway activation and drug response modelled in colorectal-tumor-derived 3D cultures. Plos Genet 15, e1008076 (2019).
This work describes how limitations in predicting the efficacy of targeted therapies based on genomic analyses can be overcome using personalized organoid models.
Pfohl U, Pflaume A, Regenbrecht M, Finkler S, Graf Ademann Q, Reinhard C, Regenbrecht CRA, Wedeken L: Precision Oncology Beyond Genomics: The Future Is Here – It is Just Not Evenly Distributed. Cells 10, 4 (2021).
Review paper showing how multi-omics technologies and appropriate in vitro models can be used to determine tumor heterogeneity and, based on this, improve clinical practice in terms of modern precision oncology.
Dahlmann M, Gambara G, Brzezicha B, Popp O, Pachmayr E, Wedeken L, Pflaume A, Mokritzkij M, Gül-Klein S, Brandl A, Schweiger-Eisbacher C, Mertins P, Hoffmann J, Keilholz U, Walther W, Regenbrecht C, Rau B, Stein U: Peritoneal metastasis of colorectal cancer (pmCRC): identification of predictive molecular signatures by a novel preclinical platform of matching pmCRC PDX/PD3D models. Molecular Cancer 20, 129 (2021).
This study is the first to report the establishment of personalized PDX/PD3D® models of peritoneal metastases from colorectal cancer and provides a thorough molecular characterization of the models.
Sankarasubramanian S, Pfohl U, Regenbrecht CRA, Reinhard C, Wedeken L: Context Matters - Why We Need to Change From a One Size Fits all Approach to Made-to-Measure Therapies for Individual Patients With Pancreatic Cancer. Front Cell Dev Biol 9, 760705 (2021).
The review discusses Reverse Clinical Engineering® as a promising method to combat lethal pancreatic cancers in personalized medicine.
Wensink GE, Elias SG, Mullenders J, Koopman M, Boj SF, Kranenburg OW, Roodhart JML: Patient-derived organoids as a predictive biomarker for treatment response in cancer patients. Npj Precis Oncol 5, 30 (2011).
This meta-analysis examined the validity and clinical utility of patient-derived tumor organoids for determining individual tumor response.
Brock A, Huang S: Precision Oncology: Between Vaguely Right and Precisely Wrong. Cancer Res 77, 23 (2017).
This review summarizes non-genetic mechanisms of heterogeneity within tumor cell populations, tumor cell plasticity, and inter-tumor cell communication that may determine the clinical outcomes of precision oncology.
Kapalczy´nska M, Kolenda T, Przybyla W, Zajaczkowska M, Teresiak A, Filas V, Ibis M, Bli´zniak R, Luczewski L, Lamparska K: 2D and 3D cell cultures - a comparison of different types of cancer cell cultures. Arch Med Sci 14, 4 (2018).
Comparison of 2D and 3D cell culture models, which are potentially more suitable for the search of novel biomarkers and new treatment strategies for personalized medicine.
Malone ER, Oliva M, Sabatini PJB, Stockley TL, Siu LL: Molecular profiling for precision cancer therapies. Genome Med 12, 1 (2020).
This review summarizes current and upcoming approaches to implementing precision cancer medicine and identifies challenges and potential solutions to facilitate the interpretation of molecular genomic analyses and maximize their clinical utility.
Collaborations












